RESEARCH
Mirror-image (D-peptide) Inhibitors
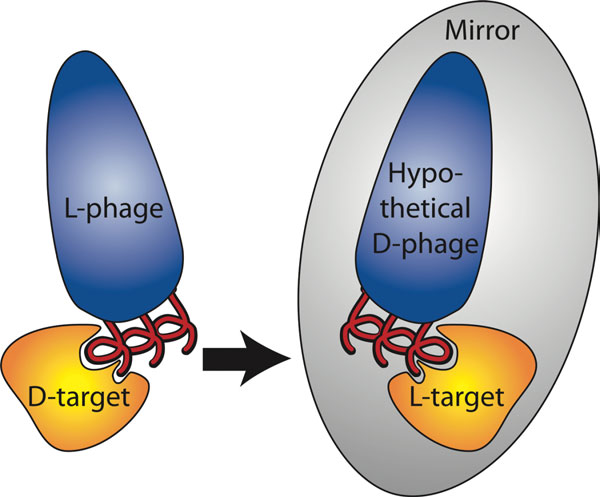
D-peptides are composed of mirror-image D-amino acids and have many potential advantages as therapeutics, including low immunogenicity and protease resistance (allowing them to last much longer in the body than natural peptides). Since there are no natural examples to guide D-peptide design, we employ a high-throughput screening method called mirror-image phage display. In traditional phage display, billions of different peptide sequences are ‘displayed’ on the bacteriophage surface (each phage bearing a different peptide). High-affinity binders are identified through several rounds of screening against a specific target. In mirror-image phage display, the target of interest is synthesized from D-amino acids to produce a mirror-image target. L-peptides displayed on phage are then selected for binding to the D-target. By symmetry, D-versions of the discovered peptides will bind to the natural L-target (see figure).
D-peptide Inhibitors of HIV Entry
HIV membrane fusion (mediated by the HIV gp41 protein) has been identified as a promising target for inhibition. Fusion is initiated by CD4/coreceptor engagement, which triggers gp41 to extend and lance the target cell before collapsing into a six-helix “trimer of hairpins” that pulls the viral and target membranes together, leading to fusion. During this conformational transition, gp41 forms a transient pre-hairpin intermediate composed of a trimeric coiled coil. We are developing D-peptide inhibitors that bind to this intermediate and prevent HIV fusion and entry. Our inhibitor design also incorporates a unique "resistance capacitor" that provides a reserve of binding energy to combat potential resistance mutations (a major problem with HIV drugs).
Our most advanced D-peptide inhibitor of HIV entry, cholesterol-conjugated PIE12-trimer, has a long half-life, potently inhibits all major HIV strains, and has a high barrier to the emergence of viral resistance. This inhibitor is also highly effective as both a preventative and therapeutic agent in the gold-standard HIV animal model and has advanced to late-stage preclinical studies with clinical trials expected to start in 2020. See figure for a model of PIE12-trimer binding to the trimeric coiled coil.
Developing D-peptides as a Broad New Therapeutic Class
We are now applying our HIV experience to other therapeutic targets ideally suited for D-peptides. We are particularly interested in using D-peptides to treat challenging antibiotic-resistant bacterial infections in the gut and urinary tract. As a highly specific inhibitor, antibacterial D-peptides have the potential to block pathogenic bacteria without disrupting the healthy microbiome. As our ability to synthesize large D-protein targets continues to improve (see below), we are now able to tackle larger cellular protein targets with applications in inflammation and cancer.
Facilitating Production of Large D-proteins
Traditionally, D-peptide discovery efforts were limited to small target proteins that can be synthesized using solid-phase peptide synthesis (SPPS) (typically <100 residues). Our lab is developing tools to create larger chemically synthesized proteins and investigating mechanisms for folding these increasingly complex proteins. Recently, we reported the chemical synthesis of a record-length (312-residue) protein, DapA. Under physiologic conditions, DapA cannot fold on its own and requires the assistance of a bacterial chaperone (GroEL/ES). Using a synthetic version of D-DapA, we showed that GroEL/ES is ‘ambidextrous’, capable of folding mirror-image DapA. This result suggests that natural chaperones can be used to fold D-proteins for mirror-image drug discovery and synthetic biology applications.
Looking ahead, we are working to make D-protein production routine by developing new tools that facilitate chemical protein synthesis, such as solubilizing “helping hands” and an automated ligator (“Aligator”) program that predicts the most efficient synthetic routes for large proteins (see figure below). Such tools are enabling us to pursue larger target proteins for mirror-image drug development. Our long-term goal is to synthesize a D-ribosome to enable rapid production of any mirror-image protein.